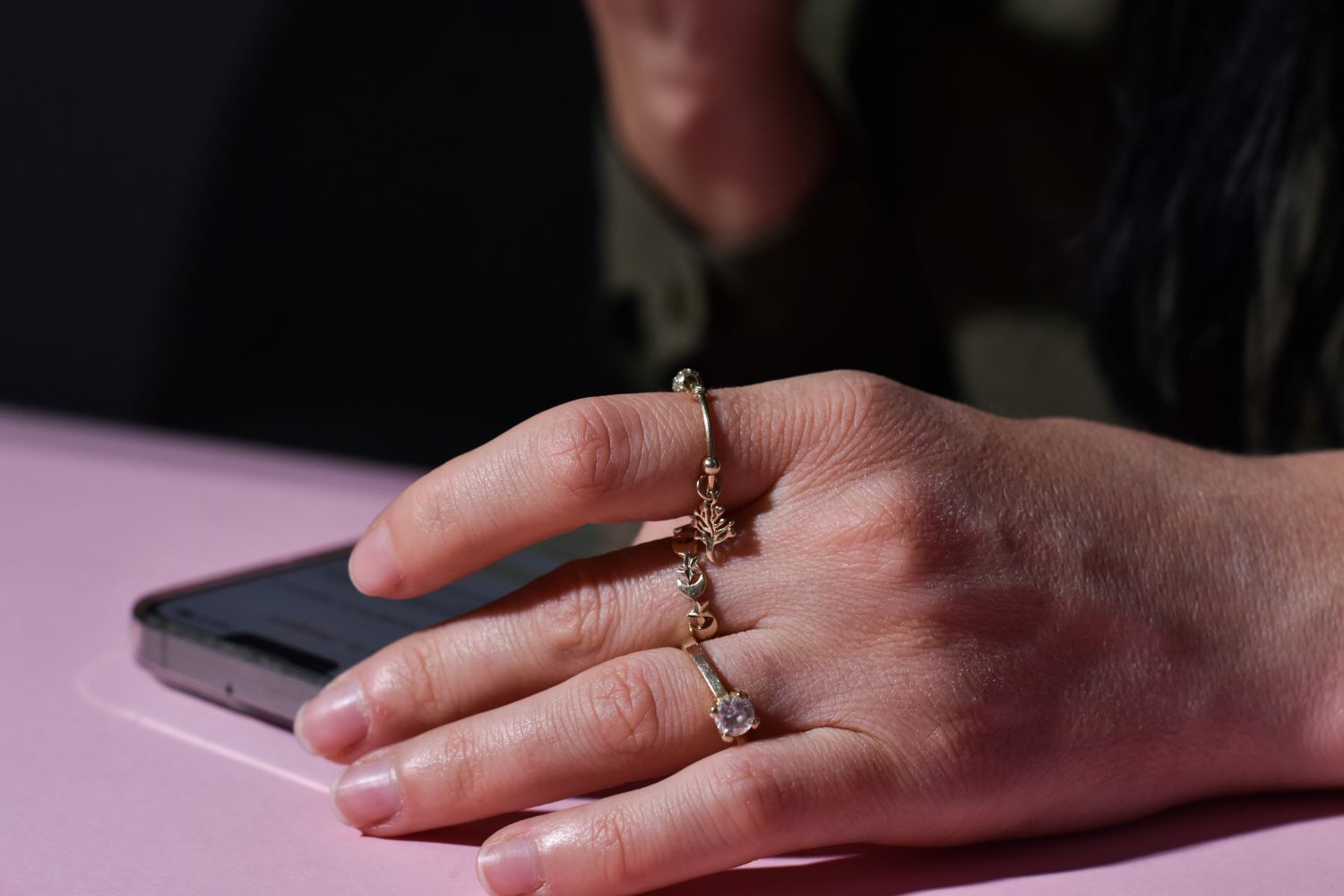
ว่ากันด้วยเรื่องของ “แหวน” เป็นอะไรที่ทุกๆ ท่าน น่าจะคิดเกี่ยวกับการเลือกแหวนหรือสวมใส่แหวนกันบ้างใช่มั้ยครับ แต่ท่านทราบเกี่ยวกับ “ความหมายเกี่ยวกับการสวมแหวน” กันบ้างหรือเปล่าครับ วันนี้เราจะขอพาทุกๆ ท่านไปพบกับ “ความหมายของการสวมแหวนแต่ละนิ้ว” กันครับ จะเป็นอย่างไรมีเนื้อหาแบบไหนนั้น…เราไปชมกันดีกว่าครับ ความหมายของการสวมแหวนบนนิ้วต่างๆ ►นิ้วโป้ง...
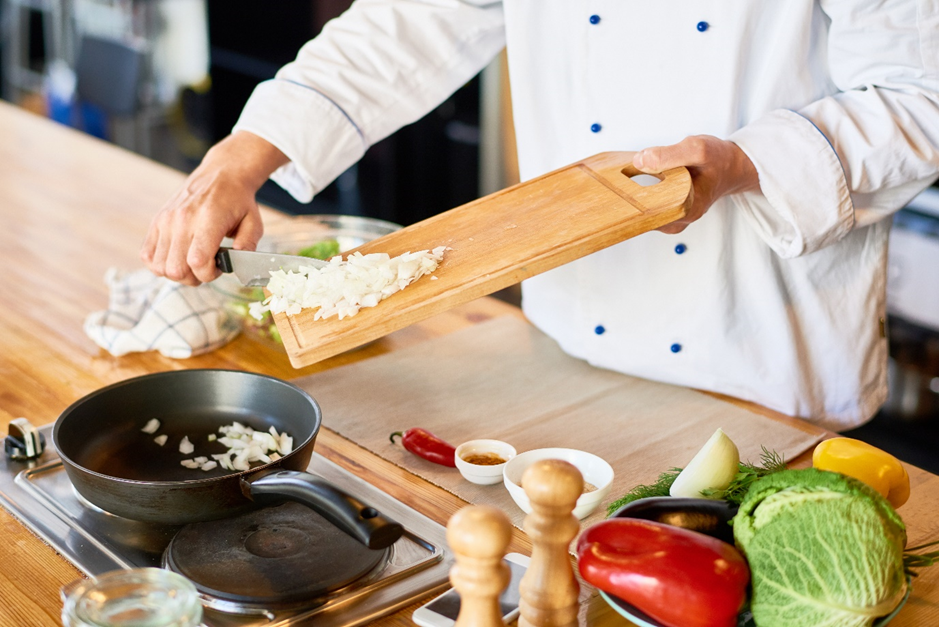
หากคุณเป็นคนรักในการทานอาหาร และอยากจะอัปสกิลเปลี่ยนจากผู้ทานมาเป็นผู้ทำดูบ้าง การหาคอร์สเรียนทำอาหารดี ๆ ที่เหมาะกับตัวเองสักคอร์สอาจไม่ใช่เรื่องง่ายนัก เพราะปัจจุบันมีคอร์สทำอาหารให้เลือกมากมาย เรียกได้ว่าเยอะมากจนเลือกไม่ถูก วันนี้เราจึงมีคู่มือในการเลือกคอร์สทำอาหารฉบับสร้างมือโปรมาฝาก ซึ่งคุณจะได้รู้ทั้งเคล็ดลับและวิธีในการเลือกคอร์สให้ถูกใจ จนตำแหน่งเชฟมือใหม่อยู่ไม่ไกลเกินเอื้อม คู่มือเลือกคอร์สเรียนทำอาหารที่ใช่ เปลี่ยนตัวเองเป็นเชฟมือโปรได้ไม่ยาก อยากเรียนทำอาหาร แต่ไม่รู้จะเริ่มจากตรงไหนก่อน...